| Journal of Medical Cases, ISSN 1923-4155 print, 1923-4163 online, Open Access |
| Article copyright, the authors; Journal compilation copyright, J Med Cases and Elmer Press Inc |
| Journal website https://jmc.elmerpub.com |
Case Report
Volume 17, Number 5, May 2026, pages 213-220
Emergence of a Citrobacter amalonaticus Strain Co-Producing Three Carbapenemases: Molecular Insights and Resistance Profiles
Hawra Alsannaa, g, Hussam Leskafia, Amani Alnimrb, c, Shouq F. Alghannamd, Sarah S. Alotaibid, Mohammad N. Alomaryd, Hatim Almutairie, Mashael Faisal Alghuraybie, Aisha Alamrif, g
aClinical Microbiology Department, Qatif Central Hospital, Al Qatif, Saudi Arabia
bLaboratory Medicine Department, King Fahad Hospital of the University, Al Khobar, Saudi Arabia
cMicrobial Genomics Lab, Department of Microbiology, College of Medicine, Imam Abdulrahman Bin Faisal University, Dammam, Saudi Arabia
dApplied Genomic Technologies Institute, King Abdulaziz City for Science and Technology (KACST), Riyadh 11442, Saudi Arabia
eNational Livestock and Fisheries Development Program (NLFDP), Riyadh, Saudi Arabia
fDepartment of Clinical Laboratory Sciences, College of Applied Medical Sciences, Imam Abdulrahman Bin Faisal University, Dammam, Saudi Arabia
gCorresponding Authors: Hawra Alsanna, Clinical Microbiology Department, Qatif Central Hospital, Al Qatif, Saudi Arabia; Aisha Alamri, Department of Clinical Laboratory Sciences, College of Applied Medical Sciences, Imam Abdulrahman Bin Faisal University, Dammam, Saudi Arabia
Manuscript submitted January 14, 2026, accepted March 3, 2026, published online March 27, 2026
Short title: Triple Carbapenemase Coproduction in C. amalonaticus
doi: https://doi.org/10.14740/jmc5285
| Abstract | ▴Top |
The emergence of carbapenemase-producing organisms poses an escalating public health risk. In this case study, we used whole-genome sequencing (WGS) to characterize the carbapenem resistance mechanisms of a multidrug-resistant Citrobacter amalonaticus (C. amalonaticus) strain recovered from a surgical site infection. The isolate was initially identified using a MALDI-TOF mass spectrometry system. Phenotypic antibiotic susceptibility testing revealed the presence of carbapenem-resistant C. amalonaticus isolate with high-level resistance to both imipenem and meropenem (imipenem minimum inhibitory concentration (MIC) ≥ 32 mg/L; meropenem MIC ≥ 32 mg/L). Rapid screening of the carbapenemase-encoding genes using the GeneXpert system revealed the presence of blaOXA-48, blaNDM, and blaKPC. WGS performed on an Illumina NovaSeq 6000 platform demonstrated the presence of blaNDM-1, which was chromosomally integrated. In silico analysis suggested that blaKPC-2 and blaOXA-48 were associated with distinct plasmid replicons (IncQ2 and IncL/M, respectively), although complete plasmid reconstruction was not possible using short-read sequencing. The convergence of multiple carbapenemase genes in C. amalonaticus indicates an urgent shift in the landscape of carbapenem resistance beyond that of traditional high-risk pathogens. These findings underscore the need for enhanced active surveillance of Citrobacter spp. as emergent reservoirs of carbapenemase genes. Early detection of these novel resistance determinants is essential for strengthening targeted infection control strategies and optimizing antimicrobial stewardship interventions at regional, national, and global levels.
Keywords: Citrobacter amalonaticus; OXA-48; NDM; KPC; Carbapenemase; Whole-genome sequencing
| Introduction | ▴Top |
Antimicrobial resistance (AMR) remains a major global concern, and prolonged exposure to antibiotics drives multidrug-resistant (MDR) infections, particularly in high-risk groups [1]. The emergence and dissemination of carbapenem-resistant pathogens are recognized as a major concern by the World Health Organization (WHO), which has ranked carbapenem-resistant Enterobacterales (CRE) as “critical priority pathogen’’ requiring urgent clinical attention [2]. In Saudi Arabia, pooled carbapenem resistance among Enterobacterales ranged from 6.6% to 18.6% [3]. However, carbapenemase production remains less common in Citrobacter spp. than in Klebsiella pneumoniae or Escherichia coli (E. coli), although emerging reports indicate rising resistance levels [4]. This emphasizes the importance of collaborative efforts and active surveillance to control the spread of these difficult-to-treat pathogens.
This report describes the genomic characterization of a carbapenem-resistant Citrobacter amalonaticus (C. amalonaticus) isolate co-harboring three carbapenemase genes (blaNDM-1, blaKPC-2, and blaOXA-48), recovered from a surgical site infection (SSI) in Eastern Saudi Arabia. To our knowledge, this is the first documented case of triple carbapenemase coproduction in C. amalonaticus in this region. The objectives of this case report are to define the complete resistance profile and genomic architecture of this isolate using whole-genome sequencing (WGS), and to contextualize this finding within the regional epidemiology of carbapenem resistance, highlighting the emerging role of Citrobacter species as potential reservoirs for high-risk carbapenemase genes.
The study was conducted in accordance with the ethical principles outlined in the Declaration of Helsinki by the World Medical Association. Ethical approval was obtained from the Institutional Review Board (IRB) at Qatif Central Hospital (Research Bioethics Committee) (Approval No. QCH-SRECO 15/2025).
| Case Report | ▴Top |
A 24-year-old male of Pakistani origin who had resided in Saudi Arabia for several years prior to admission and worked as a carpenter sustained a severe occupational injury from an electrical saw, resulting in extensive trauma to the proximal thigh involving the skin, subcutaneous tissue, muscle, femur, and major blood vessels. He was admitted in a hemodynamically unstable condition and underwent emergency vascular and multidisciplinary surgery to preserve the limb. Despite the initial salvage, progressive gangrene developed, necessitating amputation. The postoperative course was complicated by recurrent, unresolved wound infections caused by multiple organisms, the most recent of which was C. amalonaticus. This isolate was found to be carbapenem- and extensively drug-resistant; phenotypic testing demonstrated resistance to nearly all tested agents. Colistin remained reliably active (minimum inhibitory concentration (MIC) 0.94 mg/L), while amikacin, trimethoprim–sulfamethoxazole, nitrofurantoin, and fosfomycin retained activity in confirmatory testing. The isolate was subsequently used for advanced molecular characterization, including WGS.
| Material and methods | ▴Top |
Bacterial strain
C. amalonaticus was isolated from an SSI in the patient. The isolate was identified using the VITEK 2 and VITEK MS systems.
Antimicrobial susceptibility testing
Initial testing of the antibiotic sensitivity of the isolate was performed using the VITEK 2 system. The isolate was resistant to Augmentin, cefazolin, cefuroxime, ceftriaxone, ceftazidime, cefepime, piperacillin-tazobactam, meropenem, imipenem; intermediate to gentamicin; and sensitive to amikacin, trimethoprim-sulfamethoxazole, nitrofurantoin, and fosfomycin.
Further antimicrobial susceptibility testing was performed to determine the MICs of the supplemental agents. Results were as follows: colistin E-test MIC 0.94 mg/L (sensitive), ceftazidime-avibactam E-test MIC ≥ 256 mg/L (resistant), meropenem E-test MIC ≥ 32 mg/L (resistant), and imipenem E-test MIC ≥ 32 mg/L (resistant), aztreonam disc was resistant.
Molecular testing of carbapenemase-producing genes
This was carried out using the GeneXpert system (Cepheid, USA), which showed positive results for the NDM, KPC, and OXA-48 genes.
DNA extraction
An overnight culture of the Citrobacter strain was used for DNA extraction using the QIAamp DNA Kit (Qiagen, Germany) following the manufacturer’s recommendations.
Library preparation and WGS
DNA concentration was measured using a Qubit Fluorometer 2.0 to ensure accurate quantification before library preparation, using the Illumina DNA Prep kit (Illumina, USA). A total of 450 ng of genomic DNA was extracted from each sample. The DNA was fragmented using Taq buffer 1 (TB1) and tagged using bead-linked transposomes (BLT), followed by termination of the reaction using Tagment Stop Buffer. The samples were purified using Tagment Wash Buffer to remove impurities. The libraries were amplified using PCR with Enhanced PCR Mix, incorporating i5 and i7 index adapters to ensure compatibility with low-plexity pooling and maintain color balance. The amplified products were purified using Sample Purification Beads and washed using freshly prepared 80% ethanol solution. Purified libraries were resuspended in Resuspension Buffer. The library size distribution was evaluated using the LabChip® GX II Touch 24 system (Revvity, USA), which demonstrated an average fragment size of approximately 600 base pairs (bp).
Sequencing preparation and sample run on NovaSeq 6000
WGS was performed using the Illumina short-read technology, which generates high-quality paired-end reads suitable for accurate base calling, variant detection, and AMR gene identification. Libraries were pooled and adjusted to 2.7 nM as a pre-sequencing normalization step before the final dilution for loading. The denaturation process was carried out using 0.2 N sodium hydroxide (NaOH), followed by neutralization with 400 mM Tris-HCl according to the Illumina Denaturation and Neutralization Protocol. The prepared libraries were loaded onto an S2 Flow Cell (200-cycle kit) and run on a NovaSeq 6000 platform according to the manufacturer’s specifications.
Bioinformatics analysis and sequence alignment
The sequence file was subjected to comprehensive de novo genomic analysis using the Bactopia pipeline v3.2.0[5], which is built upon the nf-core framework for reproducible and scalable workflows [6]. This pipeline integrates multiple open-source tools and databases [5] to carry out quality control, genome assembly, annotation, typing, and resistance profiling.
The workflow began with raw sequencing of short reads that were quality-filtered and assembled using SPAdes v3.15.5 [7]. This step reconstructs the bacterial genome without relying on a reference, thereby enabling the study of novel or uncharacterized isolates.
Assembly quality was evaluated using contig number, total genome length, N50, L50, and GC content to indicate reasonable completeness. Gene prediction and functional annotation were performed using Prokka v1.14.6 [8] to identify both coding and non-coding genomic features. Taxonomic classification was performed using two approaches to confirm organism identity. Mash v2.3 [9] compared the assembled genome with the National Center for Biotechnology Information Reference Sequence (NCBI RefSeq) database and identified the most similar known strains. Sourmash [10] was used with the Genome Taxonomy Database (GTDB) [11] to assign standardized and phylogenetically informed taxa. Both tools provide strain-level similarities and species-level evolutionary contexts.
Strain typing was performed using multilocus sequence typing (MLST) v2.23.0, with the Citrobacter freundii scheme from the PubMLST database [12]. This approach detects allele variants in housekeeping genes to determine an isolate’s sequence type (ST), which supports epidemiological tracking and comparison across studies.
AMR and stress response genes were identified using ABRicate v1.0.1 [13], which screens genomic assemblies against curated resistance gene databases. The screened databases included the Comprehensive Antibiotic Resistance Database (CARD) [14], ResFinder [15], ARG-ANNOT [16], NCBI AMRFinderPlus [17], and BacMet [18]. Genes were reported when the alignment identity and coverage were 90%.
Virulence features, such as toxins, secretion systems, iron acquisition systems, and predicted virulence levels (pathogenicity islands and toxin clusters) were identified by screening the genome against the Virulence Factors Data Base [19] using Abricate [13]. Any coverage < 90% was excluded.
Plasmid replicon types were identified using ABRicate v1.0.1 screening against the PlasmidFinder database, applying a minimum threshold of 90% sequence identity and 90% coverage for replicon assignment. To complement replicon screening, plasmid reconstruction and contig binning were performed using MOB-suite (v3.10) via the Bactopia workflow to classify contigs into putative plasmid and chromosomal bins and to assign predicted mobility types. Putative plasmid localization of AMR genes was inferred based on 1) the co-occurrence of AMR genes and plasmid replicons on the same contig; 2) concordance of with MOB-suite plasmid bin assignments; and 3) the presence of mobilization genes. MOB-suite reconstruction relies on reference-supported clustering against curated plasmid databases, providing additional evidence for plasmid bin assignment.
Chromosomal localization of blaNDM-1 was inferred from its detection on contigs without identifiable plasmid replicon markers, lack of plasmid bin assignment, and its genomic context consistent with chromosomal regions. Given the inherent limitations of short-read sequencing, these findings represent in silico predictions of gene-plasmid associations and do not constitute definitive plasmid reconstruction. Therefore, the plasmid features and gene-plasmid associations (Tables 1, 2) represent in silico predictions rather than fully resolved plasmid structures.
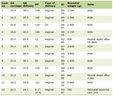 Click to view | Table 1. Predictive Features of Plasmid Replicons, Predicted Mobility (Based on the Presence of Relaxase Genes), Corresponding Accession Numbers, and the Closest Taxonomic Matches |
 Click to view | Table 2. Description of BlaOXA-48 and BlaKPC-2 Carbapenemase Gene and Its Associated Antibiotics Resistance Profile |
Results
Sequencing quality control
The sequencing run achieved a Q30 score of 88%, indicating high base-calling accuracy. Assembly quality metrics for the sample included an N50 contig length of 111,647 bp, an L50 of 14, and a sequencing depth of coverage of 99.38 ×, indicating robust genome representation.
The draft assembly, using the St. Petersburg genome de novo assembler (SPAdes), yielded 129 contigs, with a total genome size of 5,287,670 bp and a GC content of 53.34%. Functional annotation with Prokka identified 5,112 genomic features, including 5,073 coding sequences, two rRNA genes, 36 tRNA genes, and 855 hypothetical proteins, consistent with Citrobacter spp. genome.
Taxonomic classification was performed using similarity- and phylogeny-based methods. Mash analysis against the NCBI Reference Sequence (RefSeq) database identified Citrobacter species NLAE-zl-C269 (NCBI:txid 1855329) as the closest reference genome, with a Mash identity of 99.09% and 826 of 1,000 shared hashes. MLST analysis using the C. freundii scheme assigned the isolate to ST714. However, phylogenomic analysis using Sourmash and the GTDB conclusively identified the isolate as C. amalonaticus. These combined results confirmed the identity of the isolate as a member of the C. amalonaticus complex, which is similar to a known Citrobacter strain and has consistent phylogenetic placement. Given the superior resolution of genome-wide analysis and phylogenomic placement over MLST, we assigned the isolate to C. amalonaticus as per GTDB.
AMR profiling detected β-lactamase genes including blaKPC-2, blaNDM-1, blaOXA-48, each known to confer resistance to broad-spectrum and carbapenem antibiotics. Additional resistance determinants included aac(6′)-Ib for aminoglycosides, mph(A) for macrolides, qnrVC4 and oqxB for quinolones, cmlA5 for phenicols, and ble for bleomycin resistance. All genes were reported to have high nucleotide identity (≥ 96.1%) and full or near-full coverage, indicating their potential presence and functional relevance (Tables 1–3).
 Click to view | Table 3. Antimicrobial Resistance Markers Predicted to Be Chromosomally Encoded (Identified Using ResFinder, CARD, and ARG-ANNOT) |
The stress-resistance gene profile included multiple genes associated with tolerance to heavy metals, acids, and biocidal agents, such as ars (A–D), merR, merE, merA, merD, merP, merT, merC (mercury resistance), pcoA-E, pcoR, pcoS, silP, silB silA, silC, silF, silE (copper and silver resistance), in addition to qacE gene, which provides resistance to quaternary ammonium compound-based disinfectants. In silico genomic analysis suggested the presence of eight plasmid replicons, as well as integrons, transposons, and insertion sequences. Computational analysis of plasmid-associated carbapenemase genes indicated that blaNDM-1 was encoded chromosomally, whereas blaKPC-2 and blaOXA-48 appeared to be associated with the IncQ2- and IncL/M-type replicons, respectively. The high sequence similarity between these replicons and known plasmids from Klebsiella pneumoniae and K. oxytoca suggests possible horizontal gene transfer events. However, due to short-read sequencing limitations, the complete plasmid architecture, structural validation, and definitive localization of resistance genes within mobile genetic elements cannot be fully resolved. These findings should be interpreted as genomic predictions rather than as definitive plasmid reconstructions.
Nucleotide sequence accession number
The complete sequence of the strain was submitted to GenBank for a BioProject number PRJNA1284363.
| Discussion | ▴Top |
C. amalonaticus is rarely encountered in clinical infections, and its association with triple carbapenemase production has major clinical implications because carbapenemases are not the most common type of β-lactamases found in Citrobacter spp. Although most Citrobacter-related case studies focus on C. freundii, since it is a frequently isolated species, few studies have described the occurrence of MDR C. amalonaticus. For example, several reports have described the presence of multiple AMR genes in C. freundii, such as those encoding NDM, KPC, VIM, OXA-181 and OXA-48, in addition to other narrow spectrum β-lactamase-producing genes, such as the intrinsic cephalosporinase AmpC [20, 21].
In an extensive surveillance study that included 769 CRE, Cabello et al identified five isolates of C. freundii and one isolate of C. amalonaticus carrying KPC-3 and OXA-48 on different plasmid [22]. In silico analysis of our case suggested the possible occurrence of multiple plasmid replicons, of which two (IncQ2 and IncL/M) were computationally associated with the carbapenemase genes blaKPC-2 and blaOXA-48, respectively. Replicon typing and sequence similarity to known Klebsiella plasmids indicated their potential for horizontal gene transfer (Tables 1–3). The widespread occurrence of horizontal gene transfer remains one of the biggest challenges in controlling drug resistance [23].
In the present study, the presence of multiple carbapenem-inactivating genes was detected in C. amalonaticus. Upon reviewing the patient’s medical records, we noticed the development of this infection after long-term use of meropenem, which may suggest selection pressure associated with the prolonged use of carbapenems [24].
The pan-resistance of the isolate, including ceftazidime-avibactam resistance, underscores a critical therapeutic challenge. This resistance is likely due to the combined activity of the metallo-β-lactamase NDM-1 (not inhibited by avibactam) and the serine carbapenemases KPC-2 and OXA-48. This leaves only a few viable options, with in vitro activity retained only for amikacin, fosfomycin, nitrofurantoin, and colistin [25]. The potential presence of plasmid-borne blaKPC-2 and blaOXA-48 in this isolate raises serious concerns regarding infection control. Plasmids carrying carbapenemase genes facilitate horizontal gene transfer across the Enterobacterales, thereby accelerating the spread of resistance in healthcare environments. This risk is amplified in hospital settings, where selective pressure and close patient contact favor plasmid dissemination. Therefore, enhanced surveillance, strict contact precaution, and plasmid epidemiology monitoring are warranted to prevent outbreaks and limit further therapeutic compromises. In situations with limited isolation capacity, such isolates should be prioritized for strict contact isolation to prevent onward transmission [22].
To our knowledge, this is the first documented clinical C. amalonaticus isolate co-harboring three carbapenemase genes (blaNDM-1, blaKPC-2, and blaOXA-48) in Saudi Arabia. Although C. amalonaticus has been reported in local studies that evaluated the prevalence of hospital-acquired infections, no previous reports have described isolates carrying triple carbapenemase genes [26]. While our analysis identified the chromosomal localization of blaNDM-1, detailed characterization of the surrounding genetic context, including insertion sequences (e.g., ISAba125), transposons, and integrative elements, was not performed because of assembly fragmentation inherent to short-read sequencing. Chromosomal integration of carbapenemase genes is clinically significant, as it may confer greater genetic stability than plasmid carriage, potentially facilitating vertical transmission and persistence within bacterial populations [27]. The mechanism of chromosomal acquisition (e.g., plasmid-to-chromosome transfer, integrative conjugative elements, or transposon-mediated insertion) remains unresolved and warrants further investigation using long-read sequencing technologies.
This case has significant implications for AMR surveillance in Saudi Arabia and the broader Gulf region. Although carbapenem resistance in Enterobacterales has been documented regionally with pooled prevalence rates of 6.6–18.6% [3], carbapenemase production in Citrobacter spp. remains underreported. The convergence of three carbapenemase genes in a single isolate represents an alarming escalation in the complexity of resistance within the region. Given the country’s role as a hub for international healthcare and the annual Hajj season, which involves millions of visitors, the detection of such high-risk isolates underscores the need for enhanced genomic surveillance, rapid molecular diagnostics, and stringent infection prevention protocols to prevent local dissemination and cross-border transmission. This case highlights that carbapenem resistance in Saudi Arabia is no longer limited to well-recognized high-risk pathogens (Klebsiella pneumoniae, Escherichia coli) but is emerging in opportunistic genera such as Citrobacter, necessitating broadened surveillance strategies.
This case report has several limitations that warrant further investigation. Although the chromosomal localization of blaNDM-1 was identified, detailed analysis of the surrounding insertion sequences, transposons, or integrative elements was not performed; consequently, the genetic mechanism underlying chromosomal integration remains unresolved. Additionally, although the short-read sequencing approach enables comprehensive resistance profiling and robust taxonomic classification, it has inherent limitations for complete plasmid reconstruction and mobile genetic element characterization, which typically require long-read or hybrid sequencing strategies. With an average fragment size of approximately 600 bp, repetitive regions and structurally complex loci cannot be fully resolved. Genomic resolution is insufficient to definitively determine plasmid architecture, achieve complete plasmid circularization, or assign AMR genes unambiguously to plasmids versus chromosomes. Plasmid-related and localization findings should therefore be interpreted as computationally inferred rather than experimentally confirmed and are best regarded as suggestive rather than definitive.
Conclusions
Genomic characterization of this isolate revealed the co-occurrence of three high-risk carbapenemase genes (blaNDM-1, blaKPC-2, blaOXA-48) in a single C. amalonaticus strain. In silico analysis suggested that some of these resistance determinants were associated with plasmid replicons, highlighting the potential for horizontal gene transfer within Enterobacterales populations. This case highlights the urgent escalation in the epidemiology of carbapenem resistance in Saudi Arabia and underscores that such resistance is now emerging in opportunistic genera beyond classical high-risk pathogens. Collectively, these findings reinforce the need for enhanced genomic surveillance, rapid molecular diagnostics, and stringent infection prevention and control measures to prevent the dissemination of high-risk isolates.
Learning points
Carbapenem-resistant C. amalonaticus, whilst less frequently encountered than Klebsiella pneumoniae or Escherichia coli, is capable of accumulating multiple carbapenemase genes and should not be underestimated as a clinical and epidemiological threat.
Standard MIC-based susceptibility reporting does not capture the mechanistic complexity of multi-carbapenemase co-carriage; WGS is essential for characterizing resistance architecture and informing targeted infection control.
In vitro susceptibility to newer agents e.g., ceftazidime-avibactam may be rendered clinically irrelevant by the concurrent expression of multiple carbapenemases that are not inhibited by the drug.
Short-read WGS provides high-resolution resistance gene cataloguing but cannot reliably reconstruct complete plasmid topology; long-read or hybrid sequencing is required for definitive mobile genetic element characterization.
In settings with high migrant worker populations, as in the case of Gulf Cooperation Council (GCC), enhanced surveillance for carbapenemase-producing organisms in surgical and wound-care settings is warranted.
Acknowledgments
The authors would like to thank Microbiology Laboratory at Qatif Central Hospital for their assistance with laboratory testing of the isolate, and King Abdulaziz City for Science and Technology for performing the WGS work.
Financial Disclosure
No funding was received for this work.
Conflict of Interest
The authors declare no competing interests.
Informed Consent
There was no need for patient informed consent because data used were all related to laboratory finding.
Author Contributions
Conceptualization: Hawra Alsanna, Hussam Leskafi, Amani Alnimr. Investigation: Hawra Alsanna, Hussam Leskafi. Practical resources, data curation and analysis: Hawra Alsanaa, Shouq F. Alghannam, Sarah S. Alotaibi, Mohammad N. Alomary, Hatim Almutairi, Mashael Faisal Alghuraybi. Writing–original draft preparation: Hawra Alsanna, Amani Alnimr, Aisha Alamri. Visualization and supervision: Amani Alnimr, Aisha Alamri. Manuscript review and editing: all authors. All authors have read and agreed to the published version of the manuscript.
Data Availability
The data supporting the findings of this study are available from the corresponding author upon reasonable request.
Abbreviations
CP: carbapenemase-producing; SSI: surgical site infection; WGS: whole-genome sequencing; GTDB: Genome Taxonomy Database; CRE: carbapenem resistant Enterobacterales
| References | ▴Top |
- Del Rio A, Muresu N, Sotgiu G, Saderi L, Sechi I, Cossu A, Usai M, et al. High-risk clone of Klebsiella pneumoniae Co-Harbouring Class A and D Carbapenemases in Italy. Int J Environ Res Public Health. 2022;19(5):2623.
doi pubmed - Sati H, Carrara E, Savoldi A, Hansen P, Garlasco J, Campagnaro E, Boccia S, et al. The WHO Bacterial Priority Pathogens List 2024: a prioritisation study to guide research, development, and public health strategies against antimicrobial resistance. Lancet Infect Dis. 2025;25(9):1033-1043.
doi pubmed - Alshehri AA, Irekeola AA. Prevalence of carbapenem-resistant Enterobacterales (CRE) in Saudi Arabia: A systematic review and meta-analysis. Saudi Pharm J. 2024;32(11):102186.
doi pubmed - Brecher SM, Tickler IA, Tenover FC. Phenotypic and genotypic discrepancies for carbapenemase-producing Citrobacter freundii in multiple isolates from a single patient. Ann Clin Microbiol Antimicrob. 2023;22(1):24.
doi pubmed - Petit RA, 3rd, Read TD. Bactopia: a flexible pipeline for complete analysis of bacterial genomes. mSystems. 2020;5(4):e00190-e00120.
doi pubmed - Ewels PA, Peltzer A, Fillinger S, Patel H, Alneberg J, Wilm A, Garcia MU, et al. The nf-core framework for community-curated bioinformatics pipelines. Nat Biotechnol. 2020;38(3):276-278.
doi pubmed - Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19(5):455-477.
doi pubmed - Seemann T. Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068-2069.
doi pubmed - Ondov BD, Treangen TJ, Melsted P, Mallonee AB, Bergman NH, Koren S, Phillippy AM. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 2016;17(1):132.
doi pubmed - Titus Brown C, Irber L. Sourmash: a library for MinHash sketching of DNA. J Open Source Softw. 2016;1(5):27.
doi - Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A, Chaumeil PA, Hugenholtz P. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol. 2018;36(10):996-1004.
doi pubmed - Jolley KA, Maiden MC. BIGSdb: Scalable analysis of bacterial genome variation at the population level. BMC Bioinformatics. 2010;11:595.
doi pubmed - GitHub - tseemann/abricate::mag_right: mass screening of contigs for antimicrobial and virulence genes. Available from: (April 2024). [Online]. Available: https://github.com/tseemann/abricate.
- Jia B, Raphenya AR, Alcock B, Waglechner N, Guo P, Tsang KK, Lago BA, et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017;45(D1):D566-D573.
doi pubmed - Bortolaia V, Kaas RS, Ruppe E, Roberts MC, Schwarz S, Cattoir V, Philippon A, et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J Antimicrob Chemother. 2020;75(12):3491-3500.
doi pubmed - Gupta SK, Padmanabhan BR, Diene SM, Lopez-Rojas R, Kempf M, Landraud L, Rolain JM. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob Agents Chemother. 2014;58(1):212-220.
doi pubmed - Feldgarden M, Brover V, Haft DH, Prasad AB, Slotta DJ, Tolstoy I, Tyson GH, et al. Validating the AMRFinder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrob Agents Chemother. 2019;63(11):e00483-e00419.
doi pubmed - Pal C, Bengtsson-Palme J, Rensing C, Kristiansson E, Larsson DG. BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res. 2014;42(Database issue):D737-743.
doi pubmed - Chen L, Yang J, Yu J, Yao Z, Sun L, Shen Y, Jin Q. VFDB: a reference database for bacterial virulence factors. Nucleic Acids Res. 2005;33(Database issue):D325-328.
doi pubmed - Riccobono E, Salvetti S, Coppi M, Montenora I, Di Pilato V, Rossolini GM. Citrobacter freundii resistant to novel beta-lactamase inhibitor combinations and cefiderocol, co-producing class A, B and D carbapenemases encoded by transferable plasmids. J Antimicrob Chemother. 2023;78(7):1677-1682.
doi pubmed - Bedenic B, Sardelic S, Luxner J, Bosnjak Z, Varda-Brkic D, Lukic-Grlic A, Marekovic I, et al. Molecular characterization of class b carbapenemases in advanced stage of dissemination and emergence of class d carbapenemases in Enterobacteriaceae from Croatia. Infect Genet Evol. 2016;43:74-82.
doi pubmed - Cabello M, Hernandez-Garcia M, Maruri-Aransolo A, Michelena M, Perez-Viso B, Ponce-Alonso M, Canton R, et al. Occurrence of multi-carbapenemase-producing Enterobacterales in a tertiary hospital in Madrid (Spain): A new epidemiologic scenario. J Glob Antimicrob Resist. 2024;38:281-291.
doi pubmed - Kim HJ, Koo SH, Choi Q. Horizontal gene transfer of blaNDM-5 among three different enterobacteriaceae species isolated from a single patient. Clin Lab. 2024;70(8).
doi pubmed - van Loon K, Voor In 't Holt AF, Vos MC. A systematic review and meta-analyses of the clinical epidemiology of carbapenem-resistant enterobacteriaceae. Antimicrob Agents Chemother. 2018;62(1):e01730-e01717.
doi pubmed - Durante-Mangoni E, Andini R, Zampino R. Management of carbapenem-resistant Enterobacteriaceae infections. Clin Microbiol Infect. 2019;25(8):943-950.
doi pubmed - Banawas SS, Alobaidi AS, Dawoud TM, AlDehaimi A, Alsubaie FM, Abdel-Hadi A, Manikandan P. Prevalence of multidrug-resistant bacteria in healthcare-associated bloodstream infections at hospitals in Riyadh, Saudi Arabia. Pathogens. 2023;12(9):1075.
doi pubmed - Sakamoto N, Akeda Y, Sugawara Y, Matsumoto Y, Motooka D, Iida T, Hamada S. Role of Chromosome- and/or Plasmid-Located bla(NDM) on the carbapenem resistance and the gene stability in Escherichia coli. Microbiol Spectr. 2022;10(4):e0058722.
doi pubmed
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, including commercial use, provided the original work is properly cited.
Journal of Medical Cases is published by Elmer Press Inc.